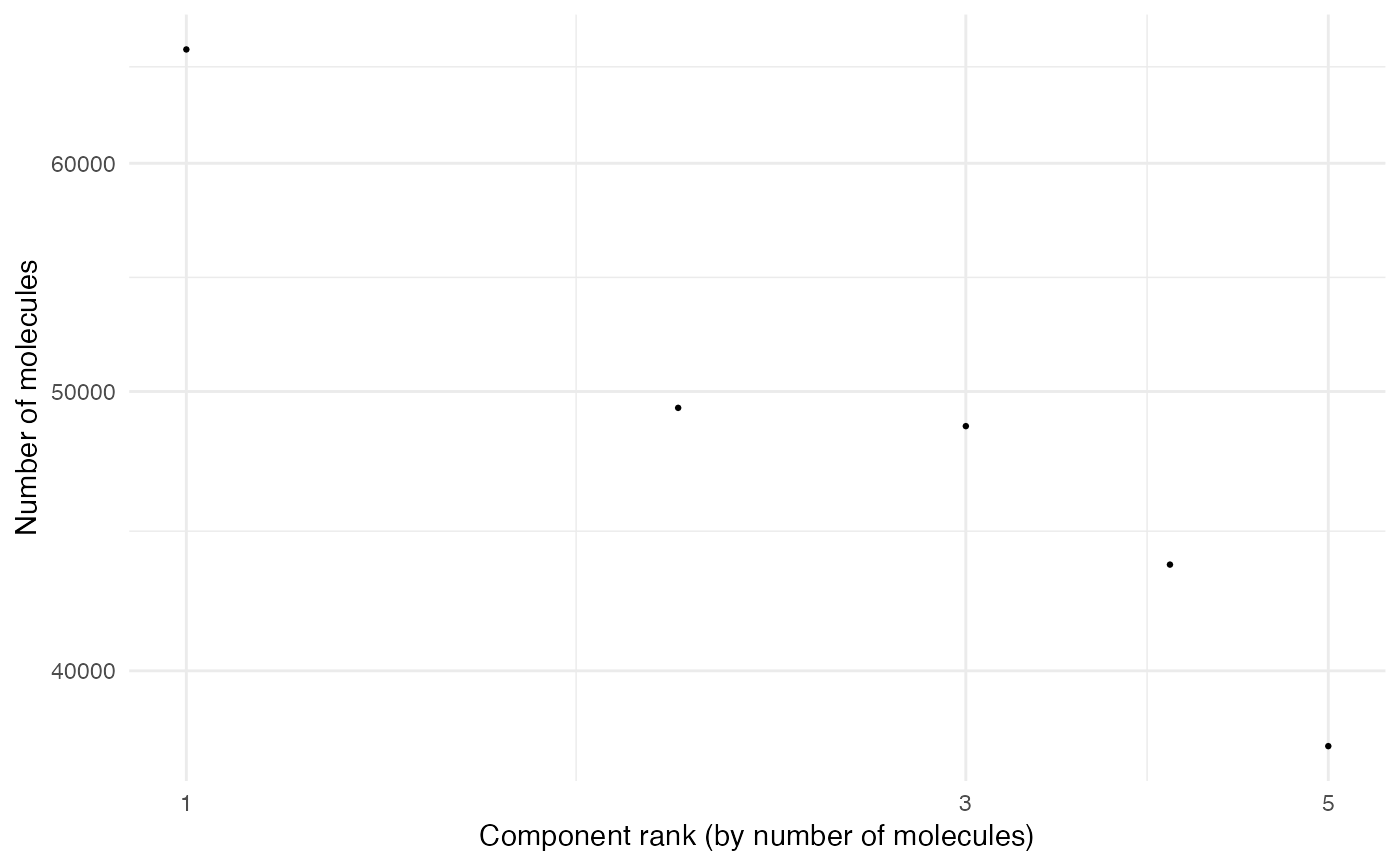
Edge Rank Plot
MoleculeRankPlot.RdPlots the number of edges/molecules per component against the component rank
Usage
MoleculeRankPlot(object, ...)
# S3 method for class 'data.frame'
MoleculeRankPlot(object, group_by = NULL, ...)
# S3 method for class 'Seurat'
MoleculeRankPlot(object, group_by = NULL, ...)Examples
library(pixelatorR)
# Load example data as a Seurat object
pxl_file_mpx <- minimal_mpx_pxl_file()
pxl_file_pna <- minimal_pna_pxl_file()
seur_obj_mpx <- ReadMPX_Seurat(pxl_file_mpx)
#> ✔ Created a 'Seurat' object with 5 cells and 80 targeted surface proteins
#> ! Failed to remove temporary file C:/Users/max/AppData/Local/Temp/RtmpmOhqBt/file62e8489454b7.h5ad
seur_obj_mpx
#> An object of class Seurat
#> 80 features across 5 samples within 1 assay
#> Active assay: mpxCells (80 features, 80 variable features)
#> 1 layer present: counts
# Plot with data.frame
MoleculeRankPlot(seur_obj_mpx[[]])
 library(pixelatorR)
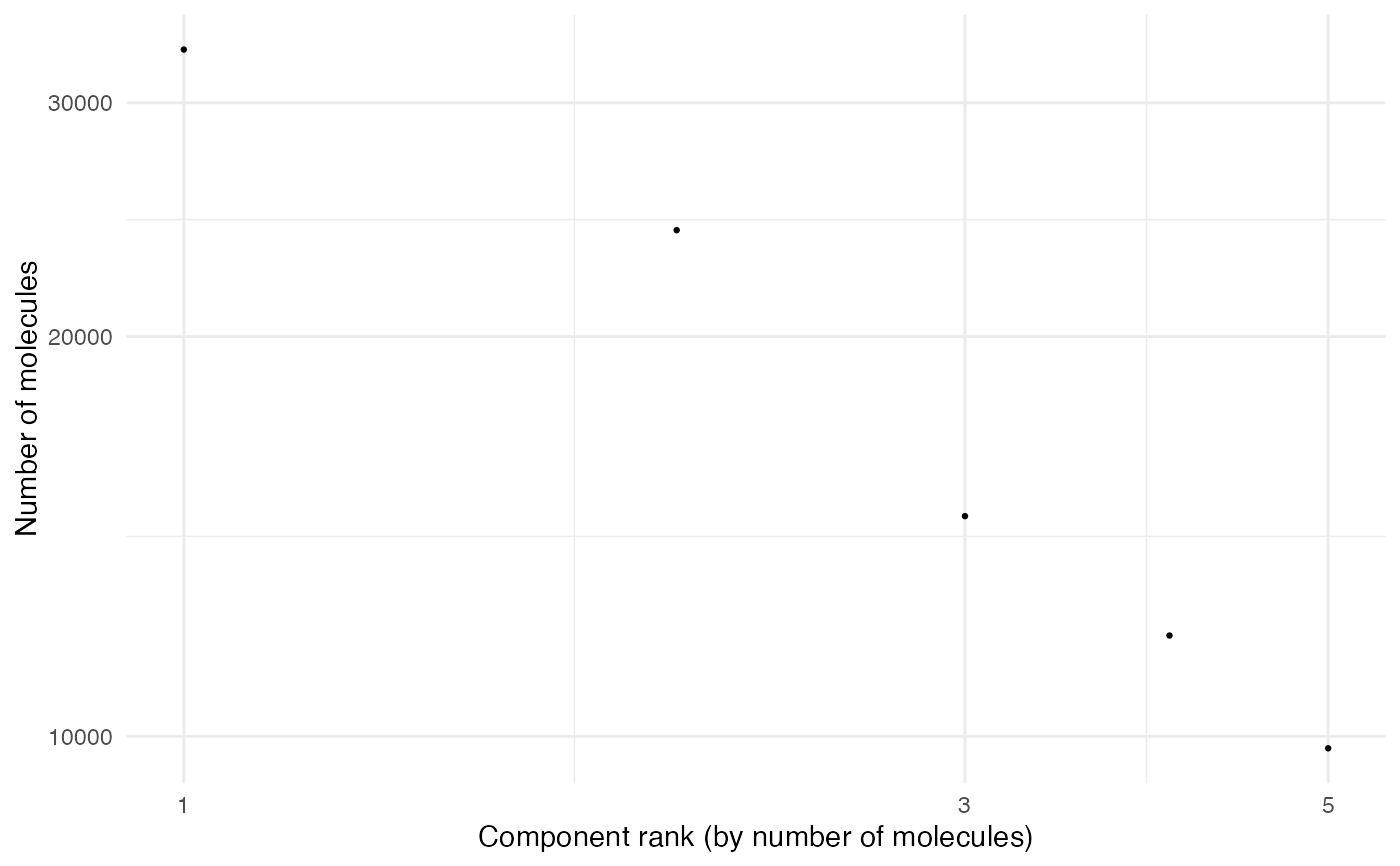
# Plot with Seurat object
MoleculeRankPlot(seur_obj_mpx)
library(pixelatorR)
# Plot with Seurat object
MoleculeRankPlot(seur_obj_mpx)
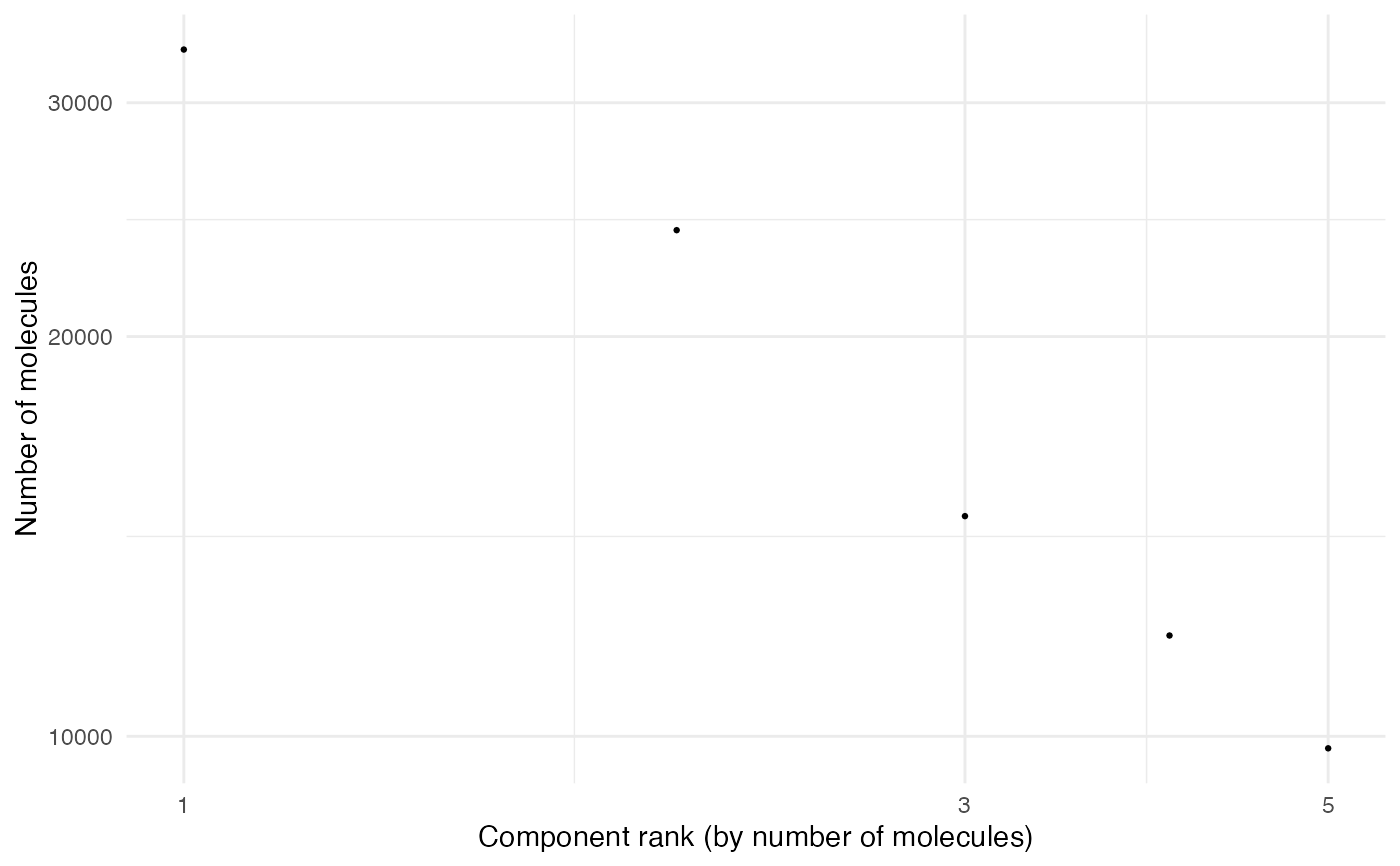 # Plot with Seurat object containing PNA data
seur_obj_pna <- ReadPNA_Seurat(pxl_file_pna)
#> ✔ Created a <Seurat> object with 5 cells and 158 targeted surface proteins
MoleculeRankPlot(seur_obj_pna)
# Plot with Seurat object containing PNA data
seur_obj_pna <- ReadPNA_Seurat(pxl_file_pna)
#> ✔ Created a <Seurat> object with 5 cells and 158 targeted surface proteins
MoleculeRankPlot(seur_obj_pna)